RSARSubsetEval is an “Attribute Selection Evaluator” algorithm (a term within Weka) that evaluates subsets of features. Evaluators for individual dataset attributes (columns/features) also exist, though naturally most data problems are defined by combinations of multiple factors (attributes), so they tend to be less applicable.
RSARSubsetEval is more widely known as a Feature Selection algorithm, which falls under the Dimensionality Reduction theme of machine learning. Essentially, it tries to reduce the number of dataset attributes (or features) your algorithm is processing. Rough Set Attribute Reduction or the “QuickReduct” algorithm is designed to quickly and measurably find a “core reduct” that leads to a consistent label across all instances. Core Reduct is a term from mathematical Rough Set Theory, a quick summary of how that works can be found in these slides. It selects those attributes that are required (according to the sample dataset instances) to distinguish the instance class labels.
Download Weka Package – RSARSubsetEval for Weka 3.7.x
- Download RSARSubsetEval Plugin – written and maintained by Peter Scully.
- Github Repo for RSARSubsetEval Plugin
- Weka on SourceForge
- Download Weka Software – v3.7.2
- Download Weka User Manual – v3.7.2
- Waikato Uni’s List of Unofficial ML packages for Weka
installATION Guide
How to install the Weka Package, via Weka’s SimpleCLI.
1. Start Weka and then Weka's simple command line interface, by clicking the 'SimpleCLI' button. 2. Paste in the following command, and press Enter: (with the path to the downloaded plugin) java weka.core.WekaPackageManager -install-package X:\path\to\weka\package\download\RSARSubsetEval.zip 3. After installation, check the package installed correctly, paste in the following, and press Enter. (this can also be checked via the Weka PackageManager) java weka.core.WekaPackageManager -package-info installed RSARSubsetEval 4. To use the package from within Weka, close the Weka software, and then re-open Weka. 5. To uninstall the package: java weka.core.WekaPackageManager -uninstall-package RSARSubsetEval
How To Use RSARSubsetEval IN a classification Pipeline:
- Open the Weka Experimenter or Weka Explorer.
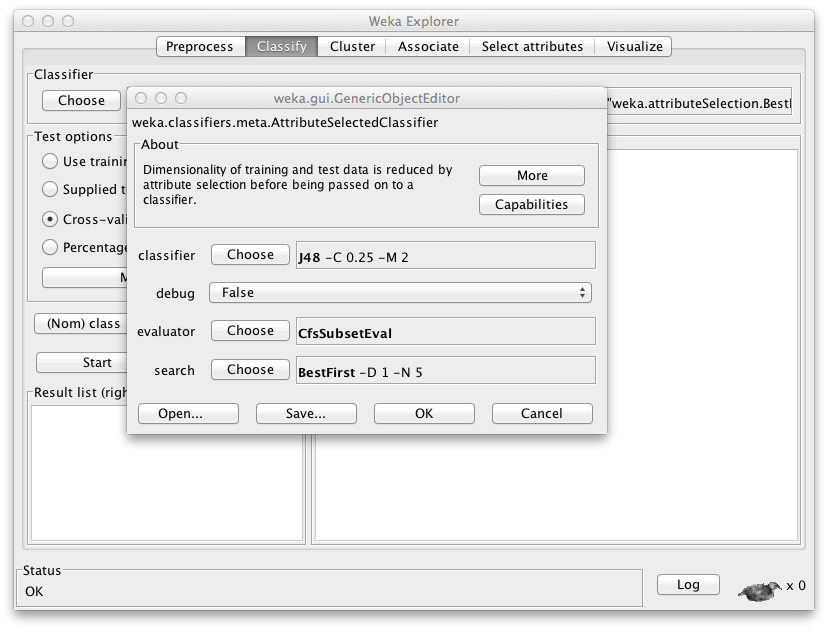
The “AttributeSelection” algorithm evaluator will be a part of a pipeline machine learning process (e.g. for model classification). - Select either AttributeSelection or AttributeSelectedClassifier.
This choice depends on the outcomes and process you want to achieve. - Choose the Evaluator as RSARSubsetEval.
If the option does not exist, follow the installation guide above (and ensure you have Weka 3.7.2 installed). - Choose the search mechanism and its parameters.
Best First Search is a good starting point.
More info on RSARSubsetEval for Weka 3.7.2
The software plugin is for the opensource data mining Weka application. The plugin was built as part of my undergraduate dissertation project at the
Department of Computer Science in Aberystwyth, Wales. The project was supervised by Dr. Richard Jensen.
The plugin algorithm is an implementation of the rough set attribute reduction (RSAR) algorithm presented by A. Chouchoulas and Q. Shen in their 2001 paper: Rough Set-Aided Keyword Reduction for Text Categorisation. This is a feature selection algorithm used to select the attributes of most importance.
The plugin was built and validated against high dimensionality gene expression datasets from the RSCTC 2010 Discovery Challenge conference competition.
The datasets tested were classified, continuous-valued, and had 25k-54k attributes with only <80 objects. Datasets were prepared from microarray experiments on patients suffering from cancers. Specifically under analysis were transcription profiles of human gliomas (E-GEOD-4290) and of gingival tissue (E-GEOD-10334).
Project
• Author: Peter Scully (pds7@aber.ac.uk)
• Supervisor: Dr. Richard Jensen
• Project: Investigating Rough Set Feature Selection for Gene Expression Analysis.
• Date: May 2011
• Location: Department of Computer Science, Aberystwyth University.
• Module: CS39440 (Undergraduate Dissertation)
• Documents: Investigating Rough Set Feature Selection for Gene Expression Analysis (BSc Dissertation PDF) • (Summary Slides PDF)
CITe
If you intend to publish your results using RSARSubsetEval in academic publications, please kindly use the following citation to reference the implementation:
Scully P.M.D., Jensen R.K (2011). Investigating rough set feature selection for gene expression analysis.(BSc Computer Science dissertation). Retrieved from (http://users.aber.ac.uk/pds7/weka/). Accessed on (xx/mm/yyyy).
or
Scully P.M.D., Jensen R.K (2011). Investigating rough set feature selection for gene expression analysis.(BSc Computer Science dissertation). Retrieved from (https://petescully.co.uk/weka/). Accessed on (xx/mm/yyyy).
Published WOrks CITING/Using Plugin
- Morán-Fernández, L., & Bolón-Canedo, V. (2023). Finding a needle in a haystack: insights on feature selection for classification tasks. In Journal of Intelligent Information Systems. Springer Science and Business Media LLC. https://doi.org/10.1007/s10844-023-00823-y
- Mkom, S. & Majid, M. A. & Sela, E.I. (2017). Performance evaluation of combined consistency-based subset evaluation and artificial neural network for recognition of dynamic Malaysian sign language. Journal of Theoretical and Applied Information Technology. 95. 2489-2496.
- Somu, N., Kirthivasan, K. & Shankar Sriram, V.S. A rough set-based hypergraph trust measure parameter selection technique for cloud service selection. J Supercomput 73, 4535–4559 (2017). https://doi.org/10.1007/s11227-017-2032-8
- Raman, M.R., Kannan, K., Pal, S.K. and Sriram, V.S., 2016. Rough Set-hypergraph-based Feature Selection Approach for Intrusion Detection Systems. Defence Science Journal, 66(6).
- Sudha et al.,Performance Comparison based on Attribute Selection Tools for Data Mining. (2014): 7(7S);61–65
- A. G. Persada, N. A. Setiawan and H. A. Nugroho, “Comparative study of attribute reduction on arrhythmia classification dataset,” 2013 International Conference on Information Technology and Electrical Engineering (ICITEE), Yogyakarta, 2013, pp. 68-72, doi: 10.1109/ICITEED.2013.6676213.
- Arafat, H., Barakat, S., & Goweda, A.F. (2012). International Journal of Emerging Trends & Technology in Computer Science ( IJETTCS ) Volume1Issue3/IJETTCS-2012-10-01-033.
Other Readings & Links
- Waikato Uni’s List of Unofficial ML packages for Weka
- RSARSubsetEval Slides on Slideplayer
- A Guide to using Attribute Selection in Weka
Thank you to the Weka team (Ebi Frank, Mark Hall, Ian Witten and the team at University of Waikato, NZ) for making this work possible.
Thanks also to Jason Brownlee for the illustrative images of applying a subset attribute evaluator to a classification pipeline.
Citations Last Updated: 26/11/23 Links Last Updated: 18/06/20. Text Last Updated: 28/08/15. First written: 25/05/11